We study the development, function, and evolution of neural circuits. We have three main interests:
1. the neuromuscular basis of behavior and how this is tuned by self-movement (proprioceptive) stimuli
2. how circuits are generated starting from neural stem cells
3. the evolution of insect motor systems
As a model, we use larval Drosophila and other immature insects. Using Drosophila allows us to span multiple levels of analysis—from molecules to behavior (see figure), and we can do so at multiple life stages. Further, we can use Drosophila as a basis to compare the biology of other immature insects.
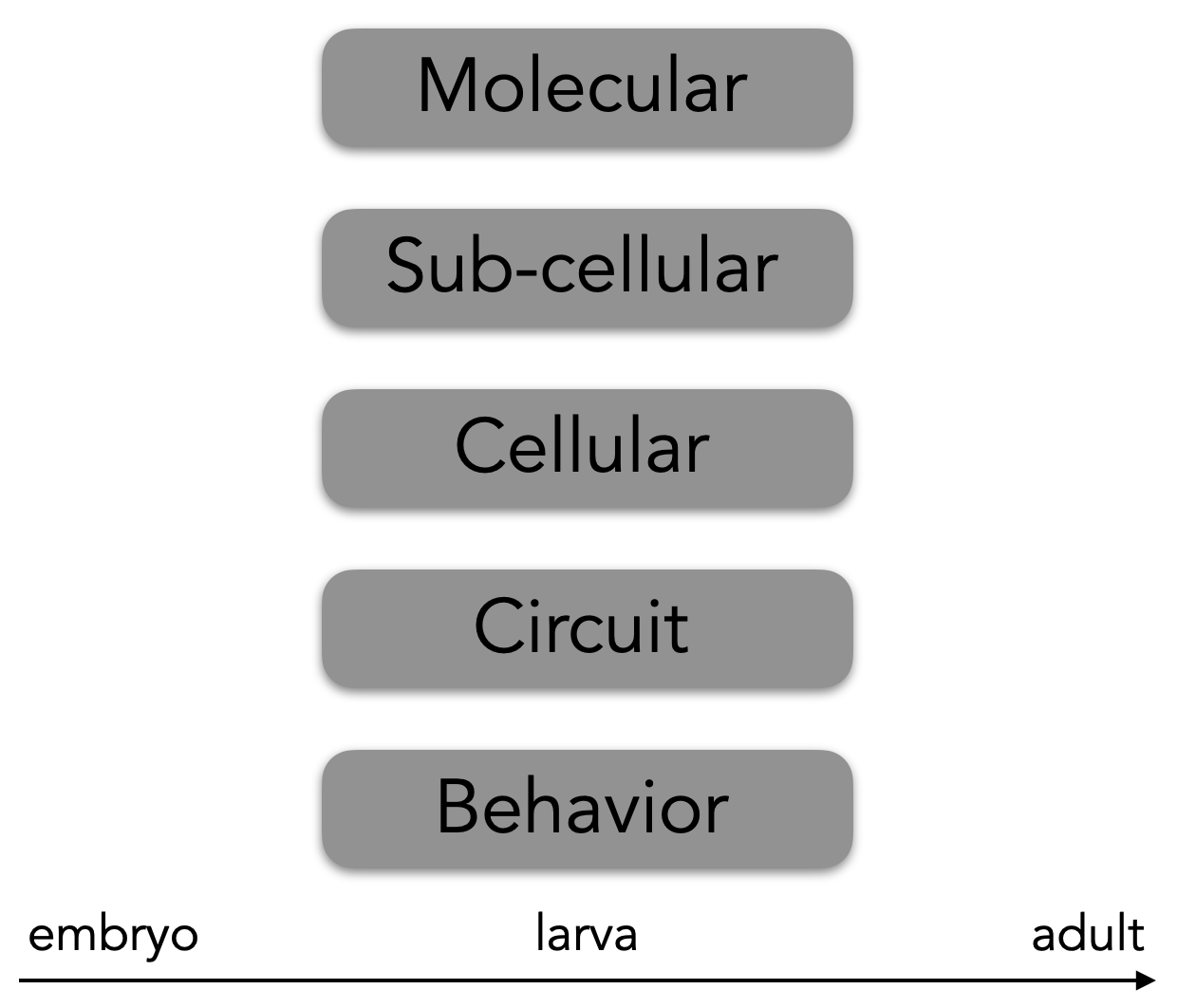
Why Drosophila larvae
As a model system, Drosophila melanogaster larvae have many experimental advantages:
Simple anatomy: The nervous system in a Drosophila larva is of “intermediate complexity” containing about 15,000 neurons. Nonetheless, we can study the Drosophila larval nervous system at single-cell resolution because the larval body and nerve cord (equivalent to the vertebrate spinal cord) is organized into repeated functional units called “hemisegments”. The number of neuromuscular cells in a single hemisegment is shown below:

Stereotyped development: The nervous system develops in a stereotyped manner with the same cells being generated in left-right hemisegments and from animal to animal.
Facile genetics: Cutting-edge molecular genetic tools allow gene manipulation in small subsets of neurons.
Optical clarity: Drosophila embryos and young larvae are optically clear, allowing us to use fluorescent microscopy to study internal events in intact organisms. For example, this allows us to use genetically encoded calcium sensors to monitor neuronal activity.
 Connectome: As a community, led by the Cardona group at the MRC Laboratory of Molecular Biology, we are in the process of building a connectome of the entire Drosophila larval nervous system. We can use this resource to identify synaptic connections between neurons and make circuit wiring diagrams.
Connectome: As a community, led by the Cardona group at the MRC Laboratory of Molecular Biology, we are in the process of building a connectome of the entire Drosophila larval nervous system. We can use this resource to identify synaptic connections between neurons and make circuit wiring diagrams.
Sensorimotor system
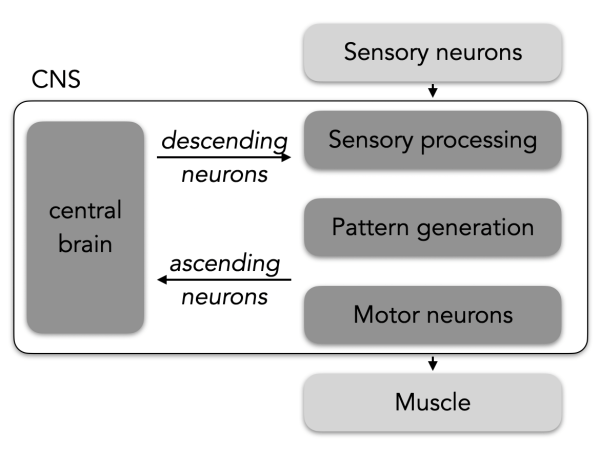
1. On the sensory side, they simultaneously process multiple different types of somatosensory stimuli (temperature, touch, self-movement). This information is used locally to tune movement or is sent to the brain, where it is used in higher-order processes like action selection. Dysfunction of these circuits can lead to devastating perceptual anomalies. For example, in allodynia, pain is caused by stimuli that do not normally provoke pain.
2. On the motor side, a network of neurons acts independently of sensory input to create patterned, rhythmic motor neuron firing. These networks are often termed “central pattern generators”. This type of circuit is unique compared to all others because it contains motor neurons, which are the only type of neuron to exit the CNS to innervate muscles and glands. When these circuits are damaged this can lead to paralysis, demonstrating their importance for all movement and behavior.
Cell types: In general, a sensorimotor circuit has at least four neuronal “functional types”. From input to output these different functional types are:
- Sensory neurons, in the peripheral nervous system, are responsible for detecting different stimuli.
- Sensory processing neurons, part of the CNS, are responsible for integrating and relaying sensory stimuli to other neurons.
- Pattern generating neurons, part of the CNS. The term “CPG” is used differently in various contexts. We use the term to mean neurons that are able to fire rhythmically in the absence of sensory input and are responsible for generating rhythmic, patterned motor neuron activity.
- Motor neurons, part of the CNS, are responsible for the control of muscle cells.
Circuit development
We seek to understand the developmental principles that lead to the self-assembly of neuronal circuits. Development of the Drosophila larval nervous system has been studied for decades and our research builds upon the many fundamental principles that have been discovered.
Evolutionarily conserved classes of interneurons: The Drosophila nerve cord contains many classes of neurons defined by expression of evolutionarily conserved, cell fate-determining transcription factors (e.g., Even-skipped [Eve]). We assess the function of each class of interneuron in control of movement and investigate the mechanistic link between molecular cell fate determination and neural circuit formation. The high conservation between vertebrate and fly nerve cords suggests that findings from the study of the Drosophila nerve cord may be directly related to the spinal cord. For example, see Marshall and Heckscher, 2022.
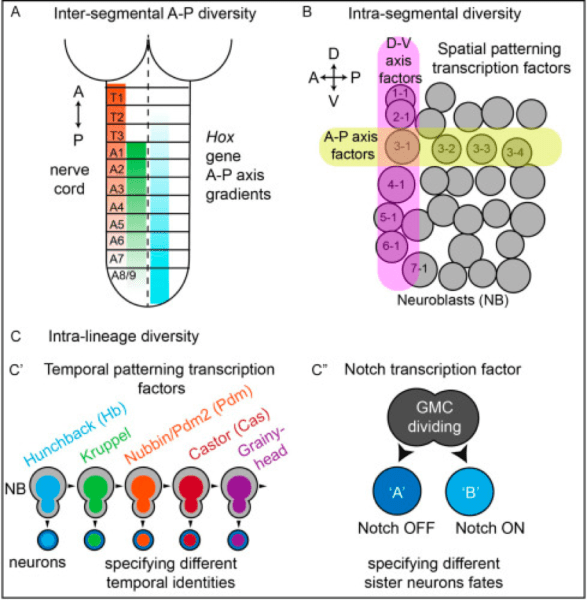
Well-characterized neuronal stem cells: During neurogenesis, a small number of neuronal stem cells (“neuroblasts” in Drosophila or [NBs]) divide to give rise to a much larger pool of diverse neuronal cell types. In general, four lineage-based mechanisms contribute to the generation of neuronal diversity. All of these developmental mechanisms operate during Drosophila neurogenesis. Specifically:
-
- Anterior-posterior diversity. The Drosophila larval CNS and larval body is segmentally organized, with three thoracic segments T1-T3, and nine abdominal segments
A1-A9. Here, Hox genes are major molecular players. - Space. Each half-segment, for example, A1 right, contains 30 neural stem cells (NB), each of which can be identified by position within a grid-like array. Molecules expressed in rows, (A-P stripes) and columns (D-V stripes) act combinatorially to make each NB unique.
- Time. Each NB generates a series of different neurons over time, always in the same birth order. Temporal transcription factors are major molecular players that specify neural temporal identity.
- Binary Cell Fate. Often, but not always, NBs give rise to a ganglion mother cell (GMC), which then divides to give rise to two neurons. These two neurons become different from each other based on Notch signaling, becoming a “Notch-ON” or “Notch-OFF” neuron.
- Anterior-posterior diversity. The Drosophila larval CNS and larval body is segmentally organized, with three thoracic segments T1-T3, and nine abdominal segments
Genetic tools: Drosophila lineage tracing and molecular genetic approaches allow us to investigate how normal and aberrant gene expression in individually identified neuronal stem cells impacts circuit assembly (see Meng and Heckscher 2019 and 2020). These experiments lead to a better understanding of neuronal circuit formation during normal development and how these processes can go awry in disease. They also have implications for the design of stem-cell-based therapies for the treatment of neuronal disease and injury.

Neuromuscular basis of behavior
How do neural circuits implement motor programs that allow animals to move? The answer to this question will provide insights relevant to neuronal basis of behavior, nervous system evolution, and are requisites for understanding principles for circuit assembly.
How do animals use muscles to move? Neurons generate movement through the control of muscles. We use live imaging of muscle contraction patterns to understand how muscle drive movement in normal animals and how these same muscle contract when subsets of neurons are perturbed.

What neurons control movement? To find neurons controlling movement we use behavioral screens, optogenetically activating small subsets of neurons and identifying behavioral defects. We then use connectomics to determine the anatomical connections among functionally important subsets of neurons. These approaches allow us to understand how microcircuits in the nerve cord control muscle contraction and locomotion, and to identify the functional architecture of the motor system at the cellular level. See Wreden et. al. 2017.
What are the processing capabilities of each set of neurons? To determine the computations performed by neurons, we precisely control input (e.g., sensory stimulation or optogenetic activation of upstream neurons). We measure the response within the neurons using calcium imaging, and monitor output with sensitive, quantitative behavioral assays. See Wang et al., 2022 eLife.
Diversity & evolution in Diptera
 Insects exploit nearly every niche imaginable—aquatic, air born, inside other animals, and more. Insects do so because they can adopt a vast array of different body plans. The most successful insects are those in which one individual adopts two body plans at different stages of the life cycle, larva and adult. For each body plan, there must be musculature and motor circuitry to control it. However, in insects, the naturally-occurring diversity in motor circuits is not well understood. The field lacks this understanding because it lacks a set of insects that are evolutionarily distant enough to be informative, yet close enough to make detailed comparisons.
Insects exploit nearly every niche imaginable—aquatic, air born, inside other animals, and more. Insects do so because they can adopt a vast array of different body plans. The most successful insects are those in which one individual adopts two body plans at different stages of the life cycle, larva and adult. For each body plan, there must be musculature and motor circuitry to control it. However, in insects, the naturally-occurring diversity in motor circuits is not well understood. The field lacks this understanding because it lacks a set of insects that are evolutionarily distant enough to be informative, yet close enough to make detailed comparisons.
Our long-term goal is to use larval insects in the Dipteran order (two-winged flies) as a system in which to understand the evolution of motor circuits and behavior. This focus is for two reasons: 1) The order spans about 250 million years of evolution, is very diverse, and contains more than 150,000 described species. Its taxonomy is well-established. Almost all Diptera undergo metamorphosis, with different larval and adult body plans that are highly adapted to their ecological niches. Thus, we will have ample species to choose from for comparative work, and we will be able to contextualize our findings with regard to habitat and evolution. 2) The dipteran order includes the well-studied model, Drosophila melanogaster. Drosophila larval locomotor behavior has been studied extensively, and Drosophila larval neuromuscular and CNS anatomy is known at single-cell resolution, including an electron-microscopic (connectome) reconstruction of all neurons and synapses. Thus, Drosophila larvae serve as a foundation for our proposed studies.
Resources
 Ellie’s YouTube channel: Content includes larval crawling data and videotutorials/protocols.
Ellie’s YouTube channel: Content includes larval crawling data and videotutorials/protocols.
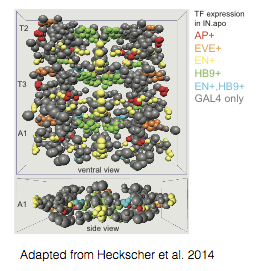
eNeuro Atlas: 3D-atlas of individual neurons in the embryonic Drosophila nerve cord that Ellie helped develop during her post-doc. This atlas established neuronal identities, molecular markers, and tools for manipulating over 60% of nerve cord interneurons. In addition, it provides a framework into which decades of work in neuro-development can be integrated (e.g., cell lineage, gene expression patterns), and has facilitated systematic analysis of the principles underlying neural circuit assembly.
